Primer sets, such as those provided by the Artic network for tiled amplicon sequencing of SARS-Cov-2, can easily be imported into Geneious from tab or comma-delimited files.
First, make a new folder in your database where you want the primers to go. We recommend using a separate folder for each set of primers. Then drag and drop the file containing your primers onto the Geneious window, and choose the format (TSV or CSV) at the prompt.
In the next window, ensure each column in the file is assigned to the correct field in Geneious, i.e the column containing the primer name is assigned to the Name field, and the sequence is in the Sequence field etc. Choose Primer as the import type, and select "Determine Characteristics" to ensure that Tm values are added to the sequence.
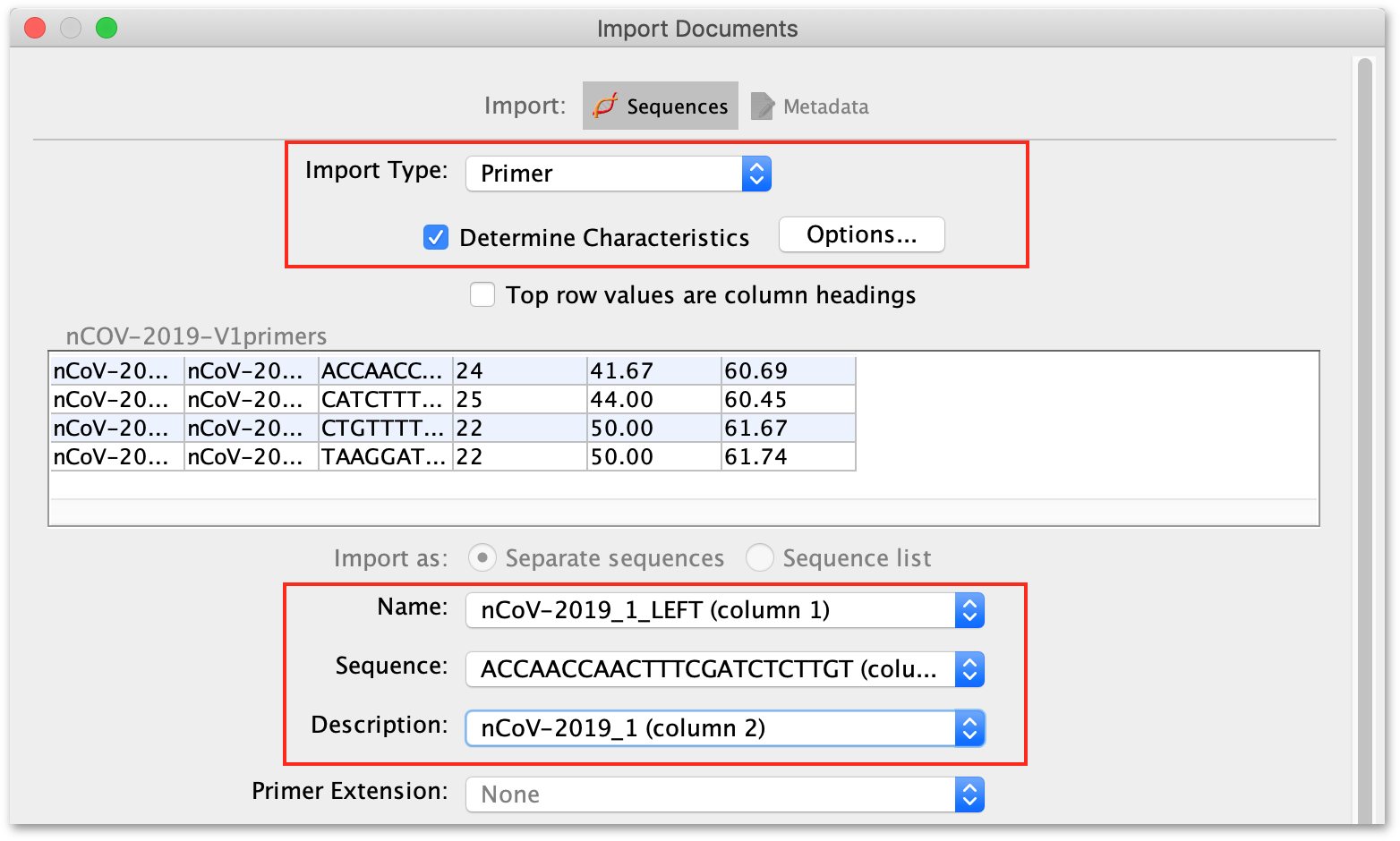
Then click OK to import the sequences.
Downloading primer sets from the Artic network
The Github repository of primer schemes is available at https://github.com/artic-network/artic-ncov2019/tree/master/primer_schemes/nCoV-2019.
Select the folder for the version you are using (e.g. V1, V2 or V3), within that folder is a .tsv file.
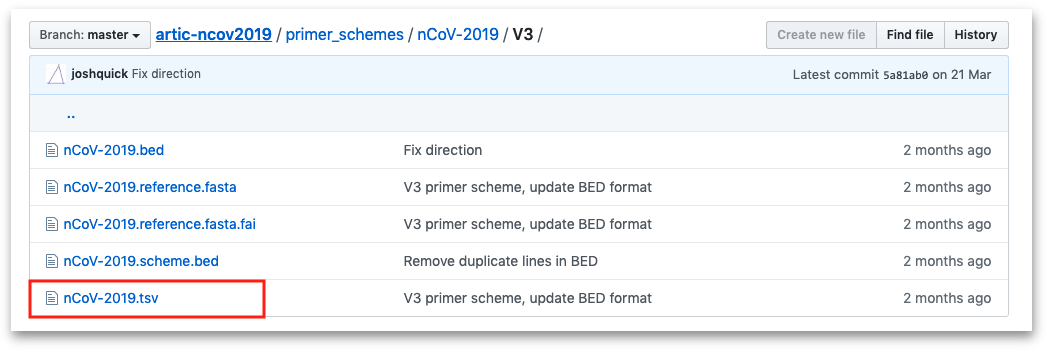
Select this file to open it, then click the "Raw" option highlighted in the screenshot below to display the file in plain text format.
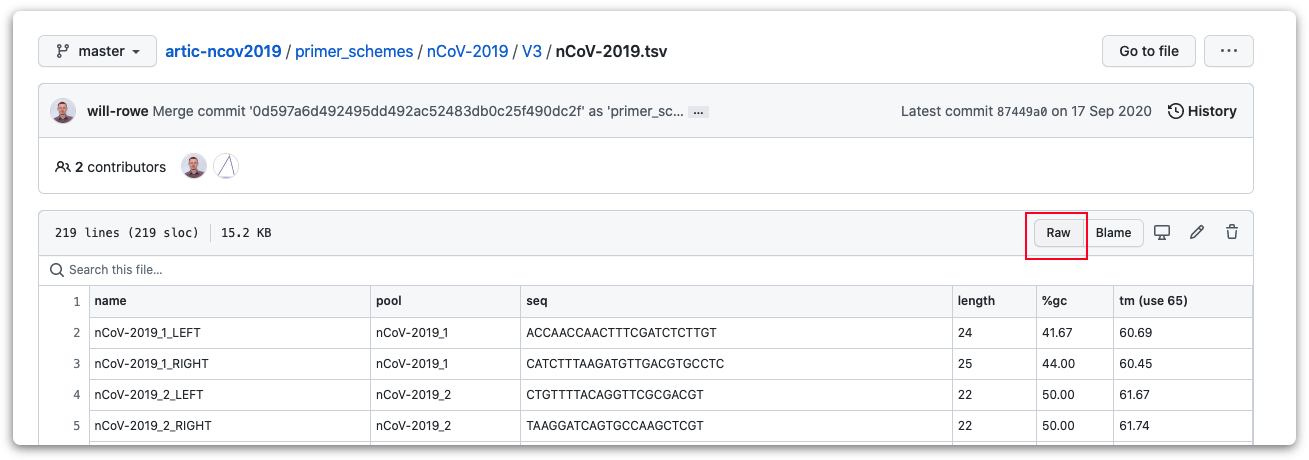
Then use control/command-A to select the contents of this file, and control/command-C to copy it.
You can then paste it directly in Geneious as a .tsv file by right-clicking in the Geneious document table and choosing "Paste" from the menu. The import will then proceed as described above and you will need to configure the columns and the import type as shown in the "Import Documents" screenshot above.